

Unlocking Antibody Potential
The success of monoclonal antibody (mAb) drugs has paved the way for next-generation antibody-based treatments, including bispecific and multispecific antibodies, antibody-drug conjugates (ADCs), and even polyclonal therapies. However, as antibody-based therapeutics develop and evolve, so does the need for comprehensive bioanalytical methods to meet regulatory requirements.
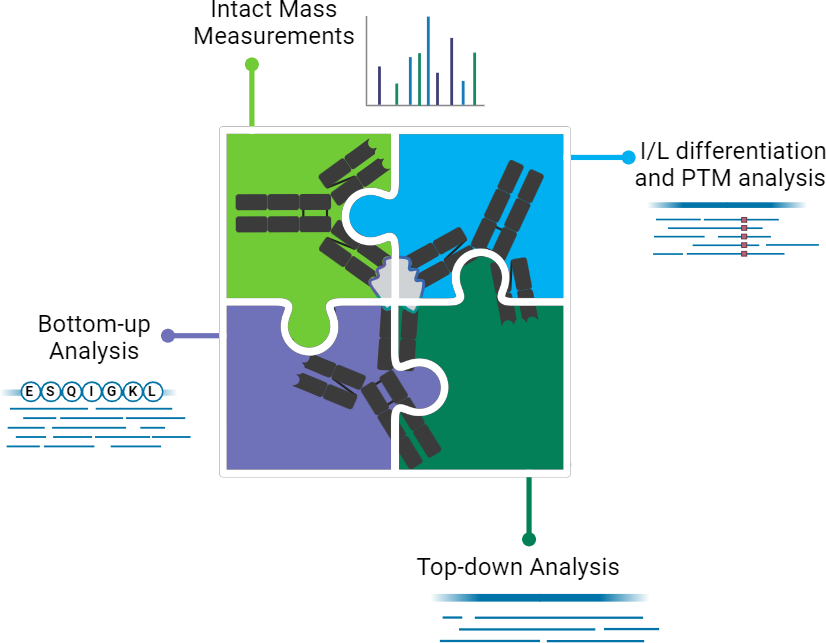
DeepAB is a powerful characterisation suite designed to support the development of antibody-derived biologics. From precise de novo sequence identification to in-depth profiling of PTMs, glycans and drug conjugation sites, our services provide the critical insights needed to optimise function, stability, and manufacturability. This platform includes:
Key Service Advantages
- Comprehensive Characterisation – Multi-level mass spectrometry workflows provide full-spectrum antibody analysis.
- AI-Powered Insights – Proprietary in-house software integrates world-leading deep learning algorithms for enhanced data interpretation.
- Advanced Sequence Resolution – Clear differentiation of isobaric residues, including 3-Tier Leu/Ile distinction and deep-learning-based retention time prediction.
Request Pricing / Schedule Consultation
Your research is important to us. We want to help you get the results you’re looking for. Contact us to schedule a free consultation to discuss your project. We will help you determine the optimal experimental setup for a fast turnaround time and with the best price.

Antibody de novo Sequencing
Key Features
- FAST: 1-2 week turnaround time
- Full and In-depth Sequence Coverage: Each amino acid is mapped to more than 20 unique peptides and 100% sequence coverage
- Accuracy: Every amino acid in CDRs is confidently supported by pairs of intense fragment ions in at least 10 MS2 scans
- Validation by Intact Mass: Intact mass measurements of heavy and light chains for double-confirmation of C-terminal lysine truncation and assembled de novo sequences
- 3-Tier Leu vs. Ile Differentiation: Differentiation of isoleucine and leucine using advanced EThcD MS method, enzyme digestion specificity and homology database analysis
- Interactive Viewer: Investigate details of the report directly from the PEAKS AB software GUI
Polyclonal Antibody de novo Sequencing
Key Features
- End-to-End Workflow Solution: Accurate sequencing of antibody mixtures directly by mass spectrometry
- Protein de novo Sequencing: No requirement for custom database generation from next-generation sequencing data
- Optimised LC: Reliable and efficient separation of simple antibody mixtures from polyclonal antibodies
- In-Depth Characterisation: Complete sequence coverage of top 4 most abundant antibody sequences
- 3-Tier Leu vs. Ile Differentiation: Differentiation of isoleucine and leucine in 4 most abundant antibodies, using advanced EThcD MS method, enzyme digestion specificity and homology database analysis
- Validation by Intact Mass: Intact mass measurements of heavy and light chains for confirmation of assembled de novo sequences for top 4 most abundant antibodies in a mixture
- PolySeq.AI: Multi-level mass spectrometry sequencing powered by innovative, deep learning-enabled data analysis algorithms
- Fast turnaround time: Comprehensive reports delivered in 4-6 weeks
ADC Characterisation
Key Features:
- Comprehensive Analysis – Multi-level mass spectrometry workflows provide a full-spectrum view of ADCs, from intact mass to peptide-level insights. Peptide mapping with high-resolution MS ensures thorough sequence verification, detecting mutations, PTM and glycan modifications.
- Advanced Conjugate Mass Analysis – Cutting-edge software algorithms enable precise identification of expected and unexpected conjugation masses.
- Accurate Drug-to-Antibody Ratio (DAR) Determination – Intact mass analysis delivers precise DAR calculations, offering insight into heterogeneity and batch-to-batch consistency.
- High-Resolution Intact & Subunit Analysis – Confirms ADC integrity and proteoform distribution, ensuring detailed characterisation of conjugated vs. unconjugated species.
Service Workflows
Service Description:
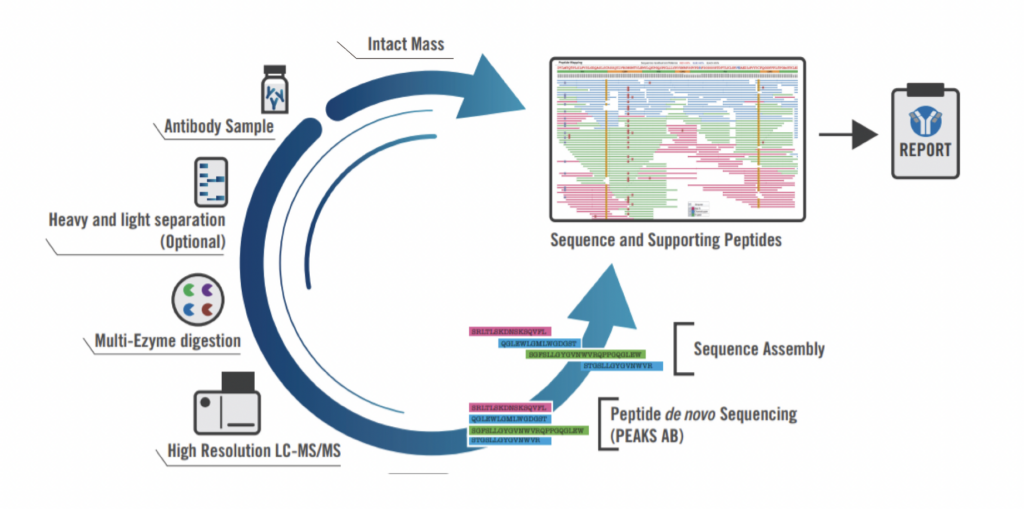
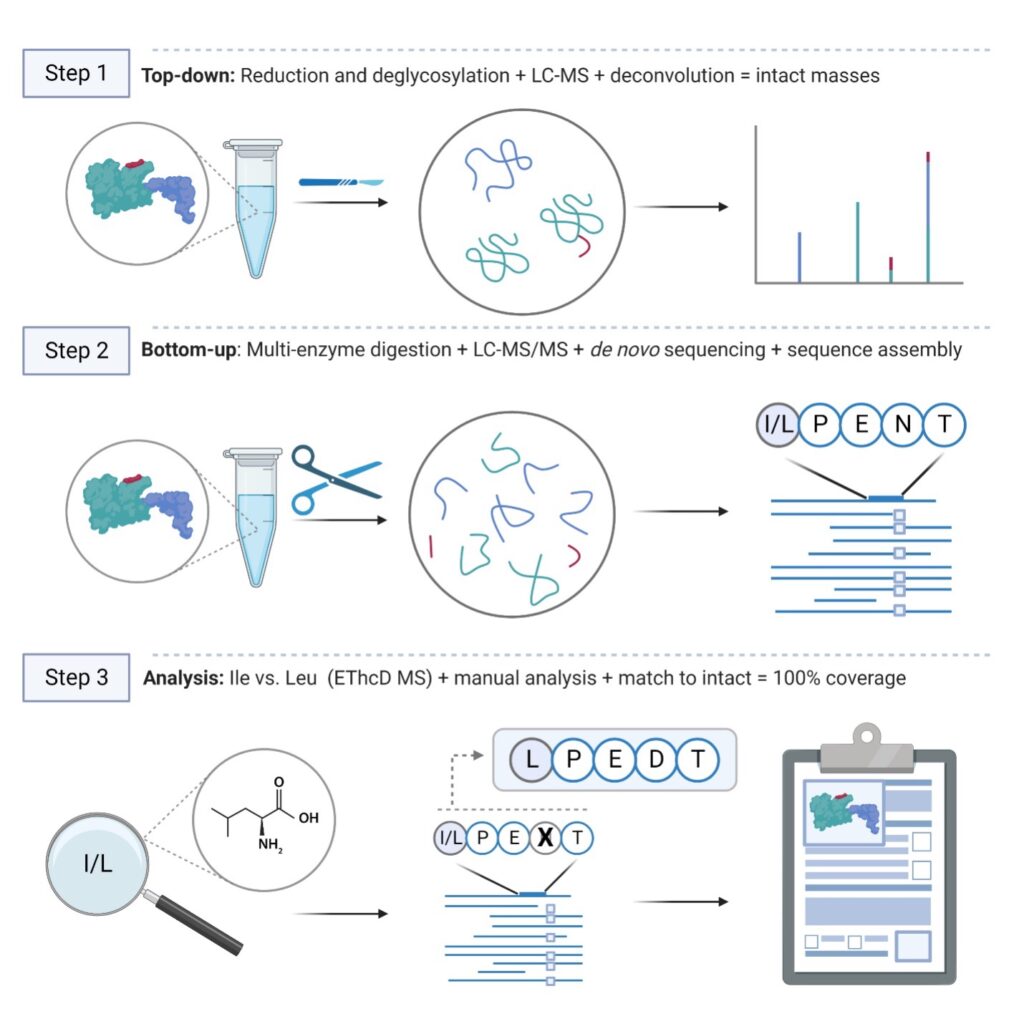
Our standard procedure integrates top-down and bottom-up MS techniques to construct and validate full protein sequences with high accuracy.
We process in-solution samples and use five different enzymes for digestion to ensure complete sequence coverage. Liquid chromatography coupled with mass spectrometry (LC-MS) generates raw peptide spectral data, which is then de novo sequenced and assembled into full protein sequences using our advanced software. Every project includes manual verification and sequence analysis to ensure accuracy. The final sequence is validated by comparing the theoretical mass to the deconvoluted intact mass. Additionally, comprehensive glycan profiling and post-translational modification (PTM) analysis are included, providing deeper insights into the structure and function of the antibody.
Workflow:
In-solution digestion → LC-MS/MS → Protein sequencing and PTM analysis
Deglycosylation → LC-MS → Intact mass deconvolution
What we need:
- For purified proteins, we require >25 μg of sample provided in-solution of in gel
- For protein mixtures (including cell lysates), a minimum of 100 μg total protein content
- Buffer information
- Target protein information eg. amino acid sequence, fasta database, species
What we deliver:
The assembly of your results and project report will include:
- Antibody sample descriptions and detailed experimental procedures
- Complete sequence of your antibody
- Intact Mass validation of reduced light and heavy chains matching the calculated average mass of the determined sequences within a ±2 Da range.
- 100% Sequence Coverage with each amino acid mapped to more than 20 unique peptides
- Spectral evidence for ambiguous amino acids
- 3-tiered differentiation of isoleucine and leucine using advanced EThcD MS method, enzyme digestion specificity and homology database analysis.
- Raw LC-MS data
Service Description:
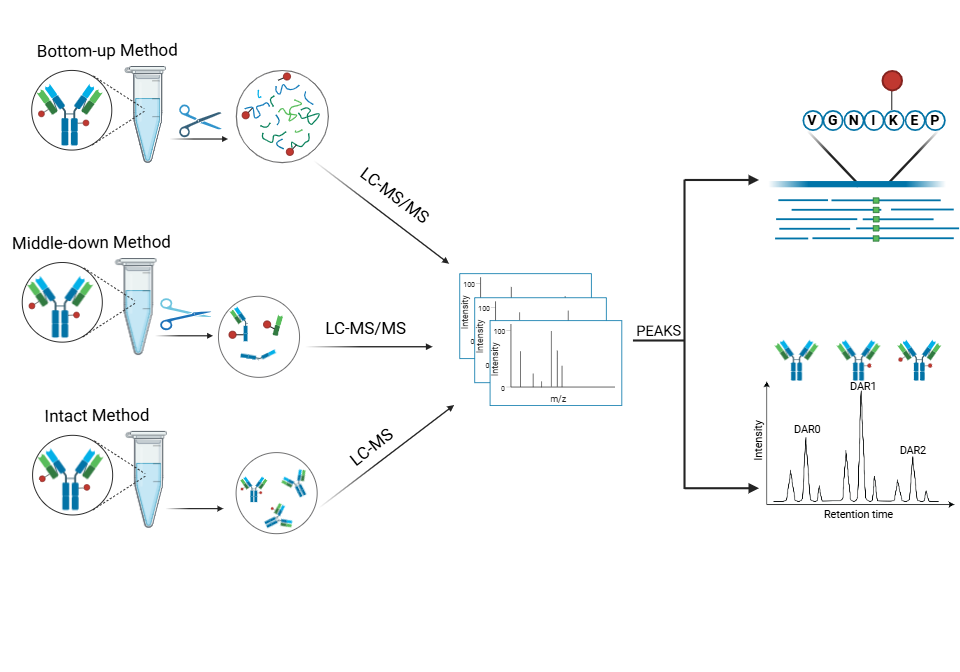
Bioanalytical methods are a critical part in antibody-drug conjugate (ADC) development for pharmacology considerations such as optimal dosing strategies and drug potency, safety, and efficacy. During early phases of research and development, this involves identifying the site(s) within the antibody where the linker and payload are conjugated, calculating drug-to-antibody ratios, and characterising the different ADC proteoforms within a sample. To address this, we optimised a complete workflow that includes sample preparation, instrumentation, and data analysis. Importantly, we integrate intact, top-down, and bottom-up mass spectrometry data for in-depth characterisation of ADC products.
Workflow:
In-solution digestion → LC-MS/MS → Antibody sequencing and PTM analysis
Deglycosylation → LC-MS → Intact mass deconvolution
What We Need:
- 20μg of sample provided in-solution or as gel bands
- Samples must be free of detergents and polymers (Triton X-100, Tween-20, PEG, etc.)
- Buffer information for in-solution samples
What We Deliver
- PEAKS ADC Report
- Antibody sequence, intact mass analysis, peptide mapping, I/L differentiation, MS2 spectra covering variable region, MS2 spectra showing conjugation site
- PEAKS Studio Viewer
- Raw data files
Method Overview
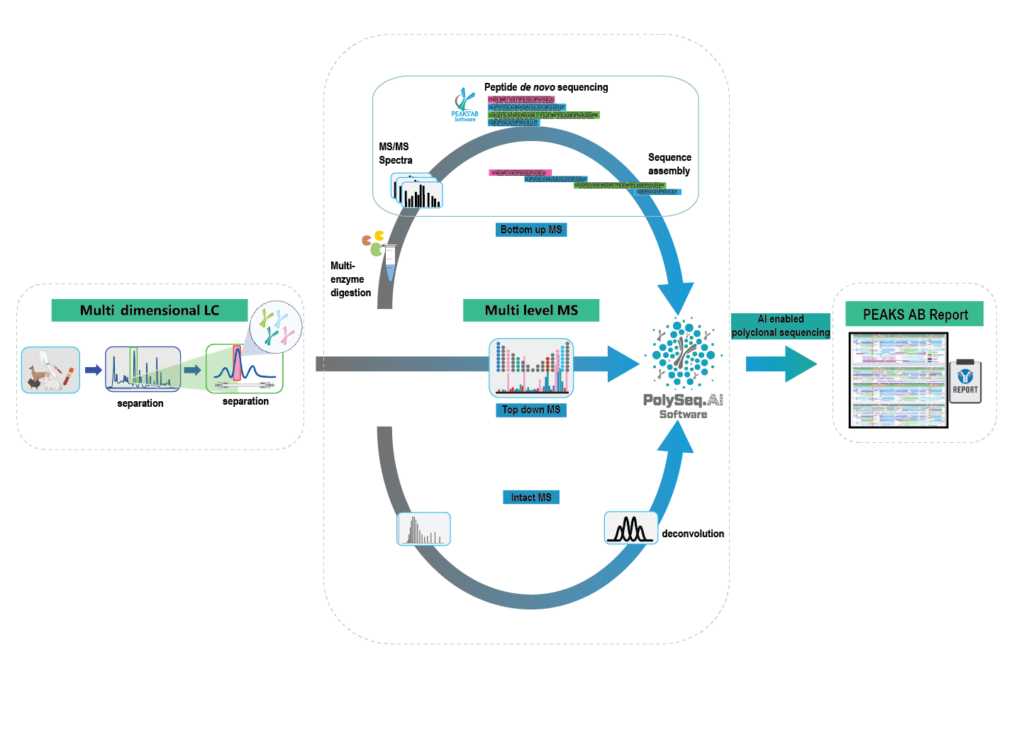
Our innovative platform combines optimised liquid chromatography (LC) separation with multi-level mass spectrometry techniques, leveraging our advanced PolySeq.AI software to resolve and accurately sequence antibody mixtures without the need for custom database generation from next-generation sequencing data. These unique capabilities, developed at BSI, build upon well-established monoclonal antibody sequencing protocols and represent a major step forward in enabling the routine sequencing of polyclonal antibodies.
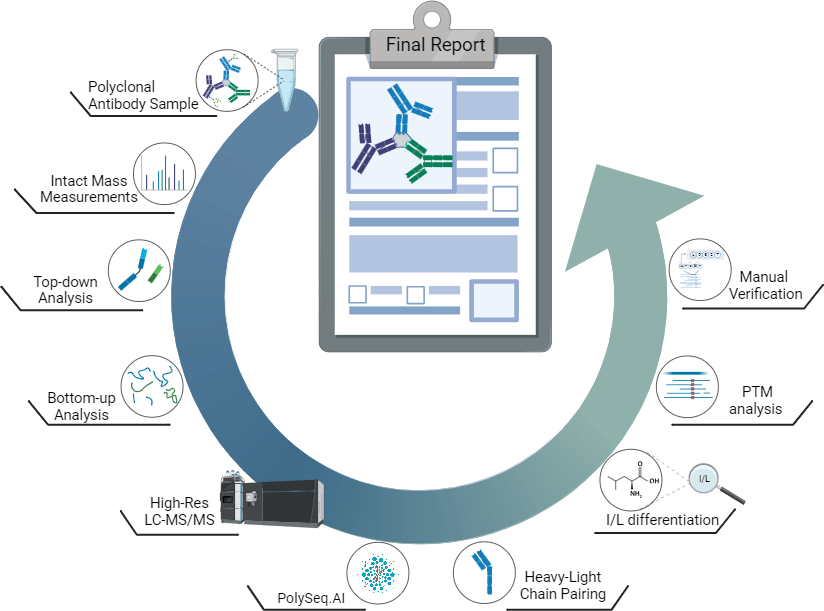
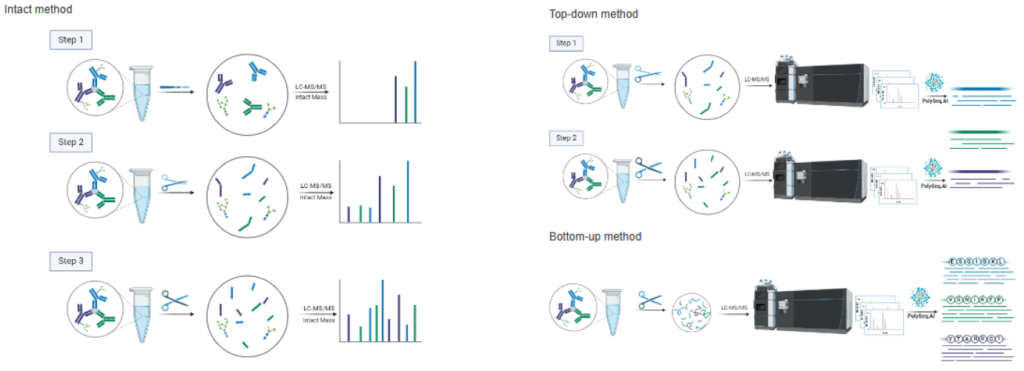
Service Description
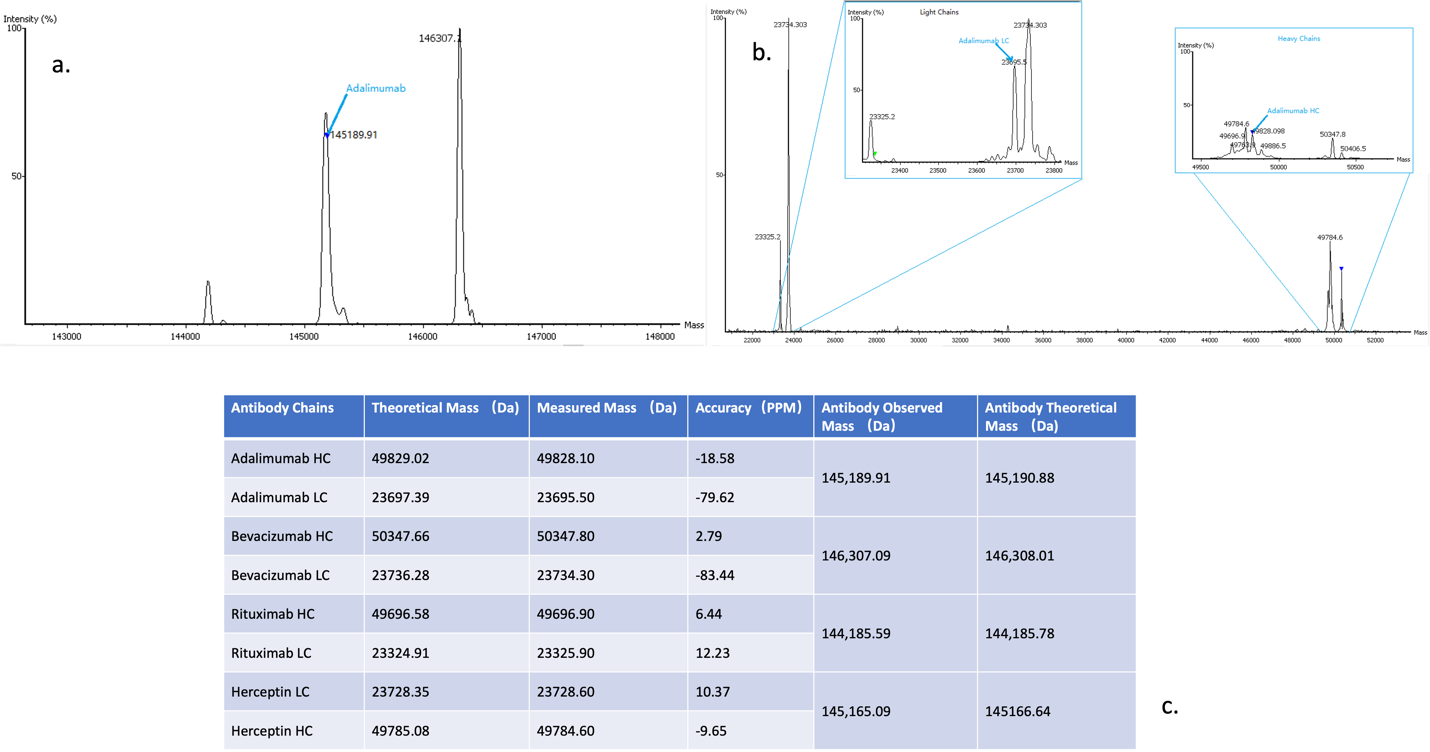
We use optimised LC methods to efficiently resolve and separate simple antibody mixtures from a complex sample. Each LC fraction is divided into 3 portions and used for intact, bottom-up, and top-down data acquisition. All data are combined within PolySeq.AI for assembly of multiple antibody sequence constructs. LC-MS and intact mass measurements are performed as illustrated below and used for correct pairing of heavy and light chains. Manual verification and sequence analysis are always included, and a comprehensive report is delivered upon project completion. Glycan profiling and a targeted search for post-translational modifications can be included as an optional add-on to this service.
What We Need:
- Sample amount: >1mg
- Samples must be free of detergents and polymers (Triton X-100, Tween-20, PEG, etc.)
- Buffer information
What We Deliver:
The assembly of your results and project report will include:
- Antibody sample descriptions and detailed experimental procedures
- Complete sequences of up to 4 antibodies per sample fraction
- Intact mass measurements with heavy and light chain pairings
- Top-down peptide mapping and fragment ion annotation
- I/L differentiation and statistics
- High-quality peptide spectra that span variable regions
- Bottom-up peptide mapping with confidence levels for each amino acid and identified post-translation modifications
References & Resources

Want to know more? Request more details to learn why our methodology and experience gives you better results than our competitors. Have questions? We’ll be happy to answer any of your inquiries today.

